AnnoTALE: Difference between revisions
| (12 intermediate revisions by the same user not shown) | |||
| Line 14: | Line 14: | ||
Jan Grau, Maik Reschke, Annett Erkes, Jana Streubel, Richard D. Morgan, Geoffrey G. Wilson, Ralf Koebnik and Jens Boch. [http://www.nature.com/articles/srep21077 AnnoTALE: bioinformatics tools for identification, annotation, and nomenclature of TALEs from ''Xanthomonas'' genomic sequences]. Scientific Reports 6:21077, DOI: 10.1038/srep21077, 2016. | Jan Grau, Maik Reschke, Annett Erkes, Jana Streubel, Richard D. Morgan, Geoffrey G. Wilson, Ralf Koebnik and Jens Boch. [http://www.nature.com/articles/srep21077 AnnoTALE: bioinformatics tools for identification, annotation, and nomenclature of TALEs from ''Xanthomonas'' genomic sequences]. Scientific Reports 6:21077, DOI: 10.1038/srep21077, 2016. | ||
For evolution-related studies using the comparative features of AnnoTALE, please also cite: | |||
Annett Erkes, Maik Reschke, Jens Boch, and Jan Grau. [https://doi.org/10.1093/gbe/evx108 Evolution of transcription activator-like effectors in Xanthomonas oryzae]. Genome Biology and Evolution, 9(6):1599–1615, 2017. | |||
If you use PrediTALE for predicting TALE targets, please also cite: | |||
Annett Erkes, Stefanie Mücke, Maik Reschke, Jens Boch, and Jan Grau. [https://doi.org/10.1371/journal.pcbi.1007206 PrediTALE: A novel model learned from quantitative data allows for new perspectives on TALE targeting]. PLOS Computational Biology, 15(7):1–31, 2019. | |||
| Line 23: | Line 32: | ||
[[File:AnnoTALEscreenshot.jpg]] | [[File:AnnoTALEscreenshot.jpg]] | ||
AnnoTALE is based on the | AnnoTALE is based on the implementation of JavaFX in Java >=8. | ||
We provide AnnoTALE as a runnable JAR file for those with a current version of Java 8 (at least update 45) on their machine. | We provide AnnoTALE as a runnable JAR file for those with a current version of Java 8 (at least update 45) on their machine. | ||
| Line 34: | Line 43: | ||
''AnnoTALE is distributed in the hope that it will be useful, but without any warranty; without even the implied warranty of merchantability or fitness for a particular purpose.'' | ''AnnoTALE is distributed in the hope that it will be useful, but without any warranty; without even the implied warranty of merchantability or fitness for a particular purpose.'' | ||
* [http://www.jstacs.de/downloads/AnnoTALE-1. | * [http://www.jstacs.de/downloads/AnnoTALE-1.5.jar Runnable Jar] (requires installed Java >= 8, update 45), may be run under Linux, macOS and Windows | ||
* | * macOS app: [http://www.jstacs.de/downloads/AnnoTALE-1.5.app-2GB.zip 2GB version], [http://www.jstacs.de/downloads/AnnoTALE-1.5.app-6GB.zip 6GB version], ZIP archive containing a macOS app including AnnoTALE and all required Java modules. For running this app, it might be required to explicitly give it running permissions in "System Preferences" -> "Security & Privacy" -> "General", which should list AnnoTALE after the first (possibly unsuccessful) starting attempt. Approve opening AnnoTALE by clicking on the button "Open Anyway" next to it. | ||
* Windows installer of AnnoTALE including Java: [http://www.jstacs.de/downloads/AnnoTALE-1. | * Windows installer of AnnoTALE including Java: [http://www.jstacs.de/downloads/AnnoTALE-1.5-2GB.exe 2GB version, 64bit Java], [http://www.jstacs.de/downloads/AnnoTALE-1.5-6GB.exe 6GB version, 64bit Java] | ||
* Windows version without installer: [http://www.jstacs.de/downloads/AnnoTALE-1.5-win.zip 6GB version, 64bit Java], ZIP archive containing AnnoTALE, all required Java modules, and a Windows batch file. For starting AnnoTALE, double-click AnnoTALE.bat. | |||
=== Source code === | === Source code === | ||
| Line 47: | Line 56: | ||
We provide an [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf AnnoTALE User Guide] in PDF format, including a detailed description of all AnnoTALE tools and installation instructions. | We provide an [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf AnnoTALE User Guide] in PDF format, including a detailed description of all AnnoTALE tools and installation instructions. | ||
== AnnoTALE command line application == | == AnnoTALE command line application == | ||
The AnnoTALE command line application is available as a [http://www.jstacs.de/downloads/AnnoTALEcli-1. | The AnnoTALE command line application is available as a [http://www.jstacs.de/downloads/AnnoTALEcli-1.5.jar runnable Jar]. For running the program and a quick help, type | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar | ||
For larger analyes, it might be necessary to increase the memory allocated by the JavaVM using the <code>-Xms</code> and <code>-Xmx</code> parameters, for instance | For larger analyes, it might be necessary to increase the memory allocated by the JavaVM using the <code>-Xms</code> and <code>-Xmx</code> parameters, for instance | ||
java -Xms512M -Xmx6G -jar AnnoTALEcli-1. | java -Xms512M -Xmx6G -jar AnnoTALEcli-1.5.jar | ||
There is no separate User Guide for the AnnoTALE command line application, but the [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf User Guide for the GUI version] describes all AnnoTALE tools, their parameters and outputs, and those of the CLI version are identical. | There is no separate User Guide for the AnnoTALE command line application, but the [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf User Guide for the GUI version] describes all AnnoTALE tools, their parameters and outputs, and those of the CLI version are identical. | ||
| Line 62: | Line 70: | ||
You obtain a list of all AnnoTALE tools by calling | You obtain a list of all AnnoTALE tools by calling | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar | ||
Output: | Output: | ||
| Line 80: | Line 88: | ||
dertale - DerTALE | dertale - DerTALE | ||
Syntax: java -jar AnnoTALEcli-1. | Syntax: java -jar AnnoTALEcli-1.5.jar <toolname> [<parameter=value> ...] | ||
Further info about the tools is given with | Further info about the tools is given with | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar <toolname> info | ||
Tool parameters are listed with | Tool parameters are listed with | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar <toolname> | ||
You get a list of input parameters by calling AnnoTALEcli-1. | You get a list of input parameters by calling AnnoTALEcli-1.5.jar with the corresponding tool name, e.g., | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar predict | ||
Output: | Output: | ||
| Line 101: | Line 109: | ||
outdir - The output directory, defaults to the current working directory (.) = . | outdir - The output directory, defaults to the current working directory (.) = . | ||
You get a description of each tool by calling AnnoTALEcli-1. | You get a description of each tool by calling AnnoTALEcli-1.5.jar with the corresponding tool name and keyword "info", e.g., | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar predict info | ||
Output: | Output: | ||
| Line 129: | Line 137: | ||
Assuming that your current working directory contains the AnnoTALEcli Jar file, a genome of interest (of a hypothetical 'Xoo' strain PXO999 with accesion CP1234567) in a FastA file "genome.fa", all rice promoters in a FastA file "Rice-promoters.fa", and a directory "out" designated to hold all output files, a typical AnnoTALE pipeline could look like | Assuming that your current working directory contains the AnnoTALEcli Jar file, a genome of interest (of a hypothetical 'Xoo' strain PXO999 with accesion CP1234567) in a FastA file "genome.fa", all rice promoters in a FastA file "Rice-promoters.fa", and a directory "out" designated to hold all output files, a typical AnnoTALE pipeline could look like | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar predict g=genome.fa outdir=out | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar analyze t=out/TALE_DNA_sequences.fasta outdir=out | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar loadAndView outdir=out | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar assign c=out/Class_builder_download.xml t=out/TALE_DNA_parts.fasta s="Xoo PXO999" a="CP1234567" outdir=out | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar rename r=out/TALE_names_\(Xoo_PXO999\).tsv i=out/Genbank__TALE_predictions.gb outdir=out | ||
java -jar AnnoTALEcli-1. | java -jar AnnoTALEcli-1.5.jar targets i=Rice-promoters.fa p="TALEs in class builder" c=out/Augmented_class_builder_\(Xoo_PXO999\).xml outdir=out | ||
Afterwards, you find all output files of all those tools in the directory "out". The output files and directories are named in analogy to the names in the AnnoTALE GUI version (see [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf User Guide for the GUI version]) | Afterwards, you find all output files of all those tools in the directory "out". The output files and directories are named in analogy to the names in the AnnoTALE GUI version (see [http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf User Guide for the GUI version]) | ||
| Line 146: | Line 154: | ||
===AnnoTALE=== | ===AnnoTALE=== | ||
'''Version 1.5''' | |||
* new "sensitive" mode of TALE Prediction tool, which may annotate TALEs in a wider range of Xanthomonas strains at the expense of an increased runtime; turned off by default | |||
* significantly improved speed of TALE Class Assignment tool | |||
* citation information for individual AnnoTALE tools available under a dedicated button in the GUI version and from the "info" command issued for individual tools in the command line version | |||
* bugfix for TALE Prediction in rather fragmented genome assemblies, where TALE predictions may extend to the ends of contigs/sequences | |||
* [http://www.jstacs.de/downloads/AnnoTALE-1.5.jar Runnable Jar] (requires installed Java >= 8, update 45), may be run under Linux, macOS and Windows | |||
* macOS app: [http://www.jstacs.de/downloads/AnnoTALE-1.5.app-2GB.zip 2GB version], [http://www.jstacs.de/downloads/AnnoTALE-1.5.app-6GB.zip 6GB version], ZIP archive containing a macOS app including AnnoTALE and all required Java modules. For running this app, it might be required to explicitly give it running permissions in "System Preferences" -> "Security & Privacy" -> "General", which should list AnnoTALE after the first (possibly unsuccessful) starting attempt. | |||
* Windows installer of AnnoTALE including Java: [http://www.jstacs.de/downloads/AnnoTALE-1.5-2GB.exe 2GB version, 64bit Java], [http://www.jstacs.de/downloads/AnnoTALE-1.5-6GB.exe 6GB version, 64bit Java] | |||
* Windows version without installer: [http://www.jstacs.de/downloads/AnnoTALE-1.5-win.zip 6GB version, 64bit Java], ZIP archive containing AnnoTALE, all required Java modules, and a Windows batch file. For starting AnnoTALE, double-click AnnoTALE.bat. | |||
'''Version 1.4.1''' | '''Version 1.4.1''' | ||
* first version to use the updated Class Builder including a large number of recently sequence strains | * first version to use the updated Class Builder including a large number of recently sequence strains | ||
| Line 210: | Line 229: | ||
=== Class Builders === | === Class Builders === | ||
* [http://www.jstacs.de/downloads/ | * [http://www.jstacs.de/downloads/class_definitions_29_07_2022.xml.gz Version 29/07/2022]: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.4.1 and later | ||
* [http://www.jstacs.de/downloads/ | * [http://www.jstacs.de/downloads/class_definitions_17_08_2021.xml.gz Version 17/08/2021]: compatible with AnnoTALE version 1.4.1 and later | ||
* [http://www.jstacs.de/downloads/ | * [http://www.jstacs.de/downloads/class_definitions_09_05_2021.xml.gz Version 09/05/2021]: compatible with AnnoTALE version 1.4.1 and later | ||
* [http://www.jstacs.de/downloads/class_definitions_10_10_2020.xml.gz Version 10/10/2020]: compatible with AnnoTALE version 1.4.1 and later | |||
* [http://www.jstacs.de/downloads/class_definitions_20_06_2019.xml.gz Version 20/06/2019]: compatible with AnnoTALE version 1.4.1 and later | |||
* [http://www.jstacs.de/downloads/class_definitions_29_09_2018.xml.gz Version 29/09/2018]: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.3 and later | |||
* [http://www.jstacs.de/downloads/class_definitions_09_03_2017.xml Version 09/03/2017]: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.2 and earlier | |||
* [http://www.jstacs.de/downloads/class_definitions_11_03_2016.xml Version 03/11/2016] | * [http://www.jstacs.de/downloads/class_definitions_11_03_2016.xml Version 03/11/2016] | ||
* [http://www.jstacs.de/downloads/class_definitions_29_01_2016.xml Version 01/29/2016] | * [http://www.jstacs.de/downloads/class_definitions_29_01_2016.xml Version 01/29/2016] | ||
* [http://www.jstacs.de/downloads/class_definitions_19_10.xml Version 10/19/2015]: used in the AnnoTALE publication (Grau ''et al.'', Sci Rep, 2016) | * [http://www.jstacs.de/downloads/class_definitions_19_10.xml Version 10/19/2015]: used in the AnnoTALE publication (Grau ''et al.'', Sci Rep, 2016) | ||
Latest revision as of 09:48, 14 September 2022
Transcription activator-like effectors (TALEs) are virulence factors of plant-pathogenic Xanthomonas spp. that function as gene activators inside plant host cells.
AnnoTALE is a suite of applications for identifying and analysing TALEs in Xanthomonas genomes, for clustering TALEs into classes by their RVD sequences, for assigning novel TALEs to existing classes, for proposing TALE names using a unified nomenclature, and for predicting targets of individual TALEs and TALE classes.
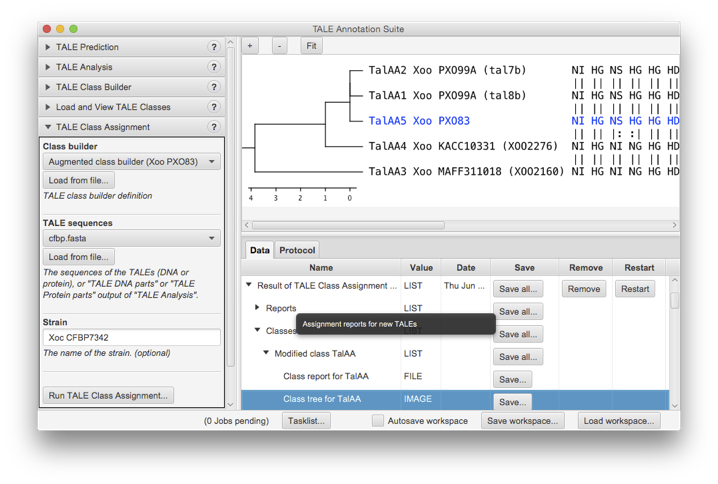
AnnoTALE is available as a JavaFX-based stand-alone application with graphical user interface for interactive analysis sessions. In addition, we provide a command line application that may be integrated into other pipelines. Both use identical code for the actual analysis, ensuring consistent results between both versions.
If you use AnnoTALE, please cite:
Jan Grau, Maik Reschke, Annett Erkes, Jana Streubel, Richard D. Morgan, Geoffrey G. Wilson, Ralf Koebnik and Jens Boch. AnnoTALE: bioinformatics tools for identification, annotation, and nomenclature of TALEs from Xanthomonas genomic sequences. Scientific Reports 6:21077, DOI: 10.1038/srep21077, 2016.
For evolution-related studies using the comparative features of AnnoTALE, please also cite:
Annett Erkes, Maik Reschke, Jens Boch, and Jan Grau. Evolution of transcription activator-like effectors in Xanthomonas oryzae. Genome Biology and Evolution, 9(6):1599–1615, 2017.
If you use PrediTALE for predicting TALE targets, please also cite:
Annett Erkes, Stefanie Mücke, Maik Reschke, Jens Boch, and Jan Grau. PrediTALE: A novel model learned from quantitative data allows for new perspectives on TALE targeting. PLOS Computational Biology, 15(7):1–31, 2019.
Important: If you would like to use the unified nomenclature of AnnoTALE in one of your publications including new TALEs or sequenced genomes, please contact us (grau@informatik.uni-halle.de) to organize the inclusion of your TALEs into the official class definition of AnnoTALE and to create stable TALE names that are unique to your TALEs.
AnnoTALE with GUI
AnnoTALE is based on the implementation of JavaFX in Java >=8.
We provide AnnoTALE as a runnable JAR file for those with a current version of Java 8 (at least update 45) on their machine.
For user's convenience, we also provide pre-packaged versions of AnnoTALE, which also include Java in the required version, for Mac OS X and Windows. Each of these versions is available two version with different memory requirements (2GB and 6GB). As long as the main memory (RAM) of your machine is sufficient, we recommend to use the 6GB version of AnnoTALE.
Download
AnnoTALE is distributed in the hope that it will be useful, but without any warranty; without even the implied warranty of merchantability or fitness for a particular purpose.
- Runnable Jar (requires installed Java >= 8, update 45), may be run under Linux, macOS and Windows
- macOS app: 2GB version, 6GB version, ZIP archive containing a macOS app including AnnoTALE and all required Java modules. For running this app, it might be required to explicitly give it running permissions in "System Preferences" -> "Security & Privacy" -> "General", which should list AnnoTALE after the first (possibly unsuccessful) starting attempt. Approve opening AnnoTALE by clicking on the button "Open Anyway" next to it.
- Windows installer of AnnoTALE including Java: 2GB version, 64bit Java, 6GB version, 64bit Java
- Windows version without installer: 6GB version, 64bit Java, ZIP archive containing AnnoTALE, all required Java modules, and a Windows batch file. For starting AnnoTALE, double-click AnnoTALE.bat.
Source code
The AnnoTALE source code is available from github.
User Guide
We provide an AnnoTALE User Guide in PDF format, including a detailed description of all AnnoTALE tools and installation instructions.
AnnoTALE command line application
The AnnoTALE command line application is available as a runnable Jar. For running the program and a quick help, type
java -jar AnnoTALEcli-1.5.jar
For larger analyes, it might be necessary to increase the memory allocated by the JavaVM using the -Xms and -Xmx parameters, for instance
java -Xms512M -Xmx6G -jar AnnoTALEcli-1.5.jar
There is no separate User Guide for the AnnoTALE command line application, but the User Guide for the GUI version describes all AnnoTALE tools, their parameters and outputs, and those of the CLI version are identical.
You obtain a list of all AnnoTALE tools by calling
java -jar AnnoTALEcli-1.5.jar
Output:
Available tools: predict - TALE Prediction analyze - TALE Analysis build - TALE Class Builder loadAndView - Load and View TALE Classes assign - TALE Class Assignment rename - Rename TALEs in File targets - Predict and Intersect Targets presence - TALE Class Presence repdiff - TALE Repeat Differences preditale - PrediTALE dertale - DerTALE Syntax: java -jar AnnoTALEcli-1.5.jar <toolname> [<parameter=value> ...] Further info about the tools is given with java -jar AnnoTALEcli-1.5.jar <toolname> info Tool parameters are listed with java -jar AnnoTALEcli-1.5.jar <toolname>
You get a list of input parameters by calling AnnoTALEcli-1.5.jar with the corresponding tool name, e.g.,
java -jar AnnoTALEcli-1.5.jar predict
Output:
At least one parameter has not been set (correctly): Parameters of tool "TALE Prediction" (predict): g - Genome (The input Xanthomonas genome in FastA or Genbank format) = null s - Strain (The name of the strain, will be used for annotated TALEs, OPTIONAL) = null outdir - The output directory, defaults to the current working directory (.) = .
You get a description of each tool by calling AnnoTALEcli-1.5.jar with the corresponding tool name and keyword "info", e.g.,
java -jar AnnoTALEcli-1.5.jar predict info
Output:
A detailed description of all tools is available in the AnnoTALE User Guide (http://www.jstacs.de/downloads/AnnoTALE-UserGuide-1.0.pdf).
*TALE Prediction* predicts transcription activator-like effector (TALE) genes in an input sequence, typically a 'Xanthomonas' genome. 'TALE Prediction' is based in HMMer nucleotide HMM models that describe N-terminus, repeat region, and C-terminus of TALEs. The input 'Genome' may be provided in FastA or Genbank format. Optionally, you may provide a strain name that will be used in the temporary TALE names and names of output files. Regardless of the input format, 'TALE Prediction' generates output in Genbank format containing the annotations of TALE genes. If the original input has already been a Genbank file, TALE annotations are added to the existing ones. In addition, 'TALE Prediction' generates annotations in GFF format, and also outputs the DNA and AS sequences of the predicted TALEs in FastA format. 'TALE Prediction' tries hard to make the CDS annotation a proper gene model, starting from a start codon and ending with a Stop. If either start or stop codon are located within the originally predicted region that is homologous to TALE genes, this original hit region is still reported as mRNA. Putative pseudo genes, e.g., with premature stop codons, are marked accordingly. The TALE DNA sequences output of 'TALE Prediction' may serve as input of the 'TALE Analysis', 'TALE Class Builder', and 'TALE Class Assignment' tools. If you experience problems using 'TALE Prediction', please contact us.
Standard pipeline
Assuming that your current working directory contains the AnnoTALEcli Jar file, a genome of interest (of a hypothetical 'Xoo' strain PXO999 with accesion CP1234567) in a FastA file "genome.fa", all rice promoters in a FastA file "Rice-promoters.fa", and a directory "out" designated to hold all output files, a typical AnnoTALE pipeline could look like
java -jar AnnoTALEcli-1.5.jar predict g=genome.fa outdir=out java -jar AnnoTALEcli-1.5.jar analyze t=out/TALE_DNA_sequences.fasta outdir=out java -jar AnnoTALEcli-1.5.jar loadAndView outdir=out java -jar AnnoTALEcli-1.5.jar assign c=out/Class_builder_download.xml t=out/TALE_DNA_parts.fasta s="Xoo PXO999" a="CP1234567" outdir=out java -jar AnnoTALEcli-1.5.jar rename r=out/TALE_names_\(Xoo_PXO999\).tsv i=out/Genbank__TALE_predictions.gb outdir=out java -jar AnnoTALEcli-1.5.jar targets i=Rice-promoters.fa p="TALEs in class builder" c=out/Augmented_class_builder_\(Xoo_PXO999\).xml outdir=out
Afterwards, you find all output files of all those tools in the directory "out". The output files and directories are named in analogy to the names in the AnnoTALE GUI version (see User Guide for the GUI version)
Version history
AnnoTALE
Version 1.5
- new "sensitive" mode of TALE Prediction tool, which may annotate TALEs in a wider range of Xanthomonas strains at the expense of an increased runtime; turned off by default
- significantly improved speed of TALE Class Assignment tool
- citation information for individual AnnoTALE tools available under a dedicated button in the GUI version and from the "info" command issued for individual tools in the command line version
- bugfix for TALE Prediction in rather fragmented genome assemblies, where TALE predictions may extend to the ends of contigs/sequences
- Runnable Jar (requires installed Java >= 8, update 45), may be run under Linux, macOS and Windows
- macOS app: 2GB version, 6GB version, ZIP archive containing a macOS app including AnnoTALE and all required Java modules. For running this app, it might be required to explicitly give it running permissions in "System Preferences" -> "Security & Privacy" -> "General", which should list AnnoTALE after the first (possibly unsuccessful) starting attempt.
- Windows installer of AnnoTALE including Java: 2GB version, 64bit Java, 6GB version, 64bit Java
- Windows version without installer: 6GB version, 64bit Java, ZIP archive containing AnnoTALE, all required Java modules, and a Windows batch file. For starting AnnoTALE, double-click AnnoTALE.bat.
Version 1.4.1
- first version to use the updated Class Builder including a large number of recently sequence strains
- minor changes to the output of the 'Load and View TALE Classes' tool, now including the accessions in the TALE sequence output
- changes to the Class Builder format to account for the increased size of class hierarchy, which previously resulted in unnecessarily large files
- 32bit/1GB Windows version no longer included
- Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE including Java: 2GB version, 6GB version
- Windows installer of AnnoTALE including Java: 2GB version, 64bit Java, 6GB version, 64bit Java
- AnnoTALE 1.4.1 command line application
Version 1.4:
- first version containing PrediTALE and DerTALE tools for target site prediction
- Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE including Java: 2GB version, 6GB version
- Windows installer of AnnoTALE including Java: 2GB version, 64bit Java, 6GB version, 64bit Java; in addition, we provide a 1GB version with 32bit Java for earlier and 32bit versions of Windows. Please use this version only if absolutely necessary, as some tools may not work due to memory restrictions.
- AnnoTALE 1.4 command line application
Version 1.3:
- AnnoTALE 1.3 Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE 1.3 including Java: AnnoTALE 1.3 2GB version, AnnoTALE 1.3 6GB version
- Windows installer of AnnoTALE 1.3 including Java: AnnoTALE 1.3 2GB version, 64bit Java, AnnoTALE 1.3 6GB version, 64bit Java; AnnoTALE 1.3 1GB version with 32bit Java
- AnnoTALE 1.3 command line application
Changes:
- modified format of Class Builder files allowing for faster download using the "Load and View TALE Classes" tool; old Class Builder files can still be loaded
- "TALE Class Presence" now also outputs a phylogenetic tree of strains based on TALEome similarities
Version 1.2:
- AnnoTALE 1.2 Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE 1.2 including Java: AnnoTALE 1.2 2GB version, AnnoTALE 1.2 6GB version
- Windows installer of AnnoTALE 1.2 including Java: AnnoTALE 1.2 2GB version, 64bit Java, AnnoTALE 1.2 6GB version, 64bit Java; AnnoTALE 1.2 1GB version with 32bit Java
- AnnoTALE 1.2 command line application
Changes:
- Results and loaded files may now be renamed in the GUI by clicking on the corresponding name in the "Data" panel
- Minor bugfixes and improvements of the GUI (Protocol may be erased, columns in "Data" panel renamed for clarity, consistency of paths in the open/save dialogs under Linux)
- Two new tools: "TALE Class Presence" and "TALE Repeat differences"
Version 1.1:
- AnnoTALE 1.1 Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE 1.1 including Java: AnnoTALE 1.1 2GB version, AnnoTALE 1.1 6GB version
- Windows installer of AnnoTALE 1.1 including Java: AnnoTALE 1.1 2GB version, 64bit Java, AnnoTALE 1.1 6GB version, 64bit Java; AnnoTALE 1.1 1GB version with 32bit Java
- AnnoTALE 1.1 command line application
Changes:
- Additional output for the "Load and View TALE Classes" tool
- "TALE Class Builder" and "TALE Class Assignment" now also accept RVD sequences (separated by dashes) as input. However, this is not recommended and some features (e.g., highlighting of aberrant repeats) will not be available. Only complete TALE DNA sequences will be accepted for inclusion into the official Class Builder.
- The internal help pages now link to the PDF User Guide
Version 1.0:
Initial AnnoTALE release
- AnnoTALE 1.0 Runnable Jar (requires Java 8, update 45 or greater)
- Mac-DMG of AnnoTALE including Java: AnnoTALE 1.0 2GB version, AnnoTALE 1.0 6GB version
- Windows installer of AnnoTALE including Java: AnnoTALE 1.0 2GB version, 64bit Java, AnnoTALE 1.0 6GB version, 64bit Java; AnnoTALE 1.0 1GB version with 32bit Java
- AnnoTALE 1.0 command line application
Class Builders
- Version 29/07/2022: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.4.1 and later
- Version 17/08/2021: compatible with AnnoTALE version 1.4.1 and later
- Version 09/05/2021: compatible with AnnoTALE version 1.4.1 and later
- Version 10/10/2020: compatible with AnnoTALE version 1.4.1 and later
- Version 20/06/2019: compatible with AnnoTALE version 1.4.1 and later
- Version 29/09/2018: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.3 and later
- Version 09/03/2017: used for "Download current definition" in "Load and View TALE Classes" within AnnoTALE version 1.2 and earlier
- Version 03/11/2016
- Version 01/29/2016
- Version 10/19/2015: used in the AnnoTALE publication (Grau et al., Sci Rep, 2016)